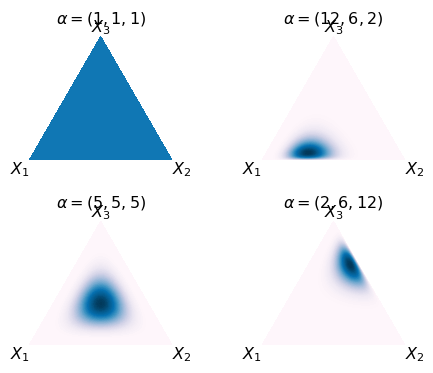Discrete Distribution Family
Multivariate discrete distribution
Intended learning outcomes
- Define the properties of Categorical, Multinomial, and Dirichlet distributions.
- Explain the use of the Dirichlet distribution as a conjugate prior and the concept of pseudo-counts.
Categorical distribution
If a random variable \(X\) has \(k\) distinct outcomes (\(k \geq 2\)), then \(X\) follows a Categorical distribution.
- Support: \(x \in \{1,\dots, k\}\)
- Parameters:
- \(p_1, p_2, \dots, p_k\) (noting that \(\sum_{i=1}^k p_i = 1\) and \(p_i \ge 0\))
- pmf
\[ p(X=i)=p_i \]
- You can view the categorical distribution as a multivariate version of the Bernoulli distribution.
- There are only \(k-1\) free parameters since \(p_k = 1 - \sum_{i=1}^{k-1} p_i\).
Example
- Phenotypes (e.g. A, B, O blood types)
- Nucleotide bases (e.g. A, T, C, G)
Multinomial distribution
Similar to the univariate case, if we summarize the counts from repeated categorical trials, we obtain the Multinomial distribution.
- Support: \(x_1,\dots,x_k \mid \sum_{i=1}^k x_i =n,\; x_i \in \mathbb{N}\)
- Parameters:
- \(p_1, p_2, \dots, p_k\) (noting \(\sum_{i=1}^k p_i = 1\) and \(p_i \ge 0\))
- \(n\): number of trials
- pmf
\[ \frac{n!}{x_1!\dots x_k!}p_1^{x_1}\dots p_k^{x_k} \]
- Again, there are only \(k-1\) free parameters since \(p_k = 1 - \sum_{i=1}^{k-1} p_i\).
Example
Summary of categorical data
Dirichlet distribution
The Dirichlet distribution is the conjugate prior of the Categorical and Multinomial distributions. It describes the belief about the probability vector \(p\) in these models.
- Support: \(p_1,\dots, p_k\)
- Parameters: \(\alpha_1,\dots, \alpha_k\)
- pdf \[ \frac{1}{B(\mathbf{\alpha})}\prod_{i=1}^{k} p_i^{\alpha_i-1} \]
- You can view the Dirichlet distribution as a multivariate version of the Beta distribution.
Bayesian estimation
- Motivation: We tries to incorporate our knowledge and experience to estimate the parameters of a distribution.
- From Bayes’ theorem, we can use the prior distribution to update our belief about the parameters after observing the data. In mathematical terms, we have
\[ p(\theta|X) = \frac{ \underbrace{p(X|\theta)}_{\text{Evidence from the data}}\cdot\underbrace{p(\theta)}_{\text{Prior belief about $\theta$}} }{\underbrace{p(X)}_{\text{Marginal likelihood of the data, not a function of $\theta$}}} \]
in short, in most of the calculation, we can write
\[ p(\theta|X) \propto p(X|\theta) \cdot p(\theta) \]
Dirichlet as categorical distribution prior
Set up
- \(X = x_1,\dots,x_N\) are \(N\) independent observations.
- Drawn from a Categorical distribution with \(\textbf{p}=(p_1,\dots,p_K)\) and \(K\) classes.
- Let \(n_k\) be the count of observations in category \(k\).
Prior
- A Dirichlet prior \(P(\textbf{p} \mid \mathbf{\alpha}) = \text{Dir}(\textbf{p} \mid \mathbf{\alpha}) = \frac{1}{\text{B}(\mathbf{\alpha})} \prod_{i=1}^K p_i^{\alpha_i - 1}\)
From Bayes’ theorem, we have
\[ \small P(\textbf{p} \mid X) \propto P(X \mid \textbf{p}) P(\mathbf{p} \mid \mathbf{\alpha}) \]
For simplification, let \(c=(c_1,\dots,c_k)\) as the count of observations in each category, and the posterior distribution of the parameter is
\[ \small \begin{aligned} P(\textbf{p} \mid \textbf{X} ) \propto & \left( \prod_{i=1}^K p_i^{c_i} \right) \left( \prod_{i=1}^K p_i^{\alpha_i - 1} \right) \\ \propto & \prod_{i=1}^K p_i^{c_i + \alpha_i - 1} \end{aligned} \]
The Posterior Predictive Distribution
Thep posterior distribution has the kernel (functional form) of a Dirichlet distribution with updated parameters \(\mathbf{\alpha}'\):
\[ \small \mathbf{\alpha}' = (\alpha_1 + c_1, \alpha_2 + c_2, \dots, \alpha_K + c_K) \]
Interpretation: adding \(\alpha_i\) pseudo-counts to the observed counts \(c_i\) for each category \(i\).
Posterior predictive distribution
Intuitively speaking if we want to guess the probability of the next observation \(x_{N+1}\) being category \(j\) is still a Categorical distribution,
but we need to use the updated parameters \(\mathbf{\alpha}'\) to calculate the :
\[ P(x_{N+1} = j \mid D, \mathbf{\alpha}) = \frac{\alpha_j + c_j}{\sum_{i=1}^K (\alpha_i + c_i)} \]
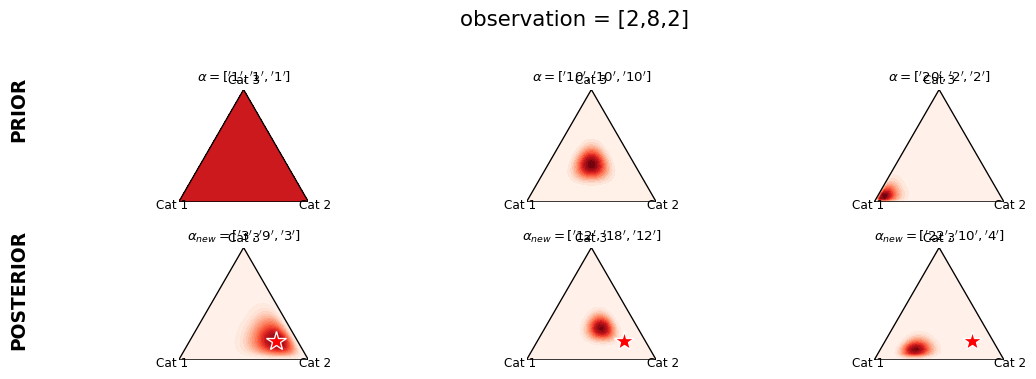
Illustration of Dirichlet Prior
Dirichlet prior adds a psuedo \(\alpha\) observations. (The observed data is [8,2,2])
Dirichlet as a multinomial distribution prior
Set up
- \(X = x_1,\dots,x_n\) are observations drawn from a Multinomial distribution with \(P=(p_1,\dots,p_K)\) and \(K\) classes.
Prior
A Dirichlet prior \(P(\textbf{p} \mid \mathbf{\alpha}) = \text{Dir}(\textbf{p} \mid \mathbf{\alpha}) = \frac{1}{\text{B}(\mathbf{\alpha})} \prod_{i=1}^K p_i^{\alpha_i - 1}\)
The probability of observing this data is
\[ \small p(X|p)= \frac{n!}{x_1!\dots x_k!}\prod_{j=1}^K p_j^{x_j} \propto \prod_{j=1}^K p_j^{x_j} \]
Using Bayes’ Rule, the posterior is proportional to the Likelihood times the Prior: \[ P(\textbf{p} \mid \textbf{X}, \mathbf{\alpha}) \propto P(\textbf{X} \mid \textbf{p}) P(\textbf{p} \mid \mathbf{\alpha}) \]
Dropping the constants (terms not involving \(\textbf{p}\)) to find the kernel:
\[ P(\textbf{p} \mid \mathbf{x}, \mathbf{\alpha}) \propto \left( \prod_{i=1}^K p_i^{x_i} \right) \left( \prod_{i=1}^K p_i^{\alpha_i - 1} \right) \]
Illustration
Thep posterior distribution has the kernel (functional form) of a Dirichlet distribution with updated parameters \(\mathbf{\alpha}'\):
\[ \begin{aligned} \mathbf{\alpha}' = (\mathbf{\alpha} + \textbf{x}) = (\alpha_1 + x_1, \alpha_2 + x_2, \dots, \alpha_K + x_K) \end{aligned} \]
Interpretation: The \(\alpha_i\) values act as “imaginary” observations seen before the experiment.
Interpretation of Dirichlet prior
| Categorical | Multinomial | |
|---|---|---|
| Prior | Dirichlet | Dirichlet |
| Posterior | \(\prod_{k=1}^Kp_k^{n_k+\alpha_k-1}\) | \(\prod_{k=1}^Kp_k^{x_k+\alpha_k-1}\) |
| Posterior predictive | Categorical | Dirichlet-multinomial |
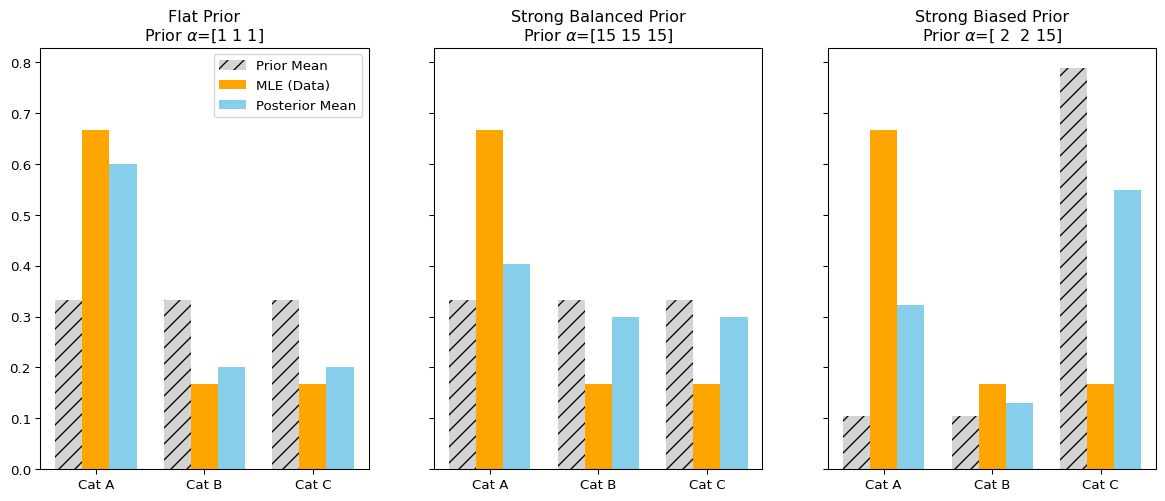
- Interpretation:
- Adding a Dirichlet prior is equivalent to adding \(\alpha\) pseudo-observations to the data.
- As the number of observations increases, the influence of the prior decreases.
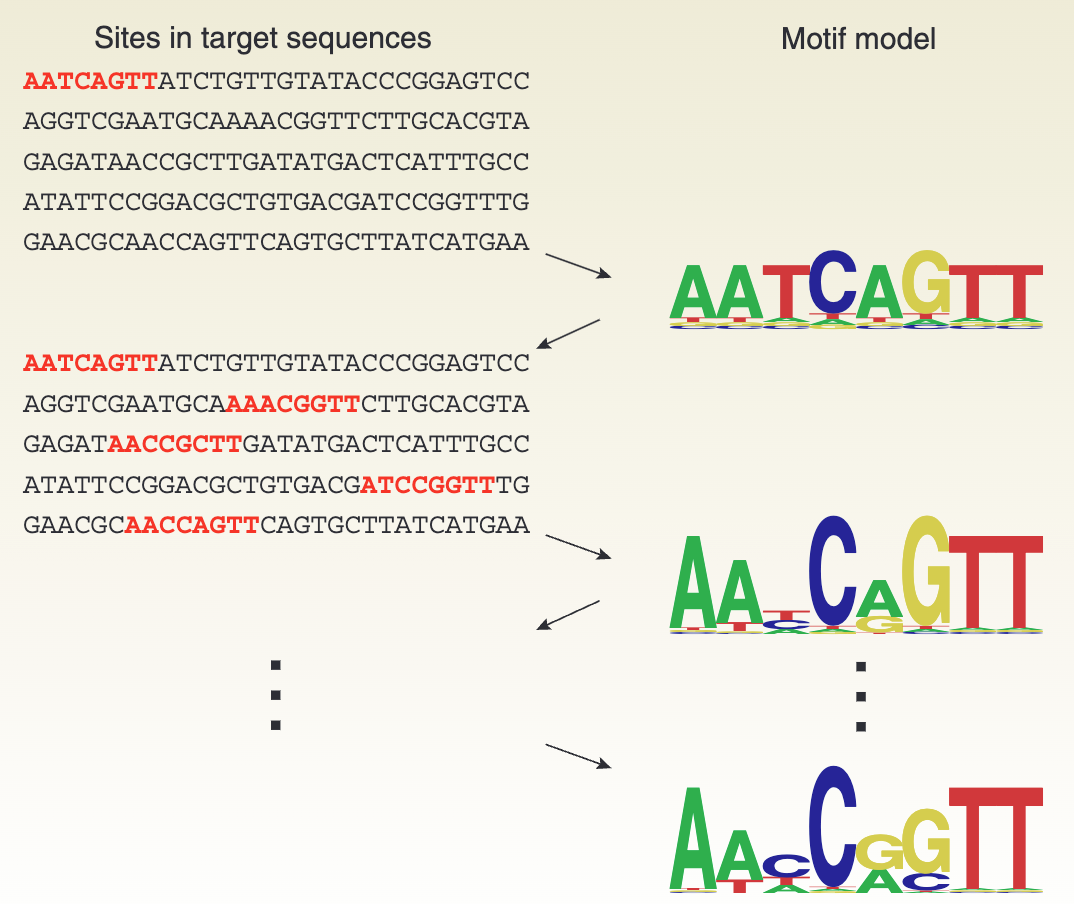
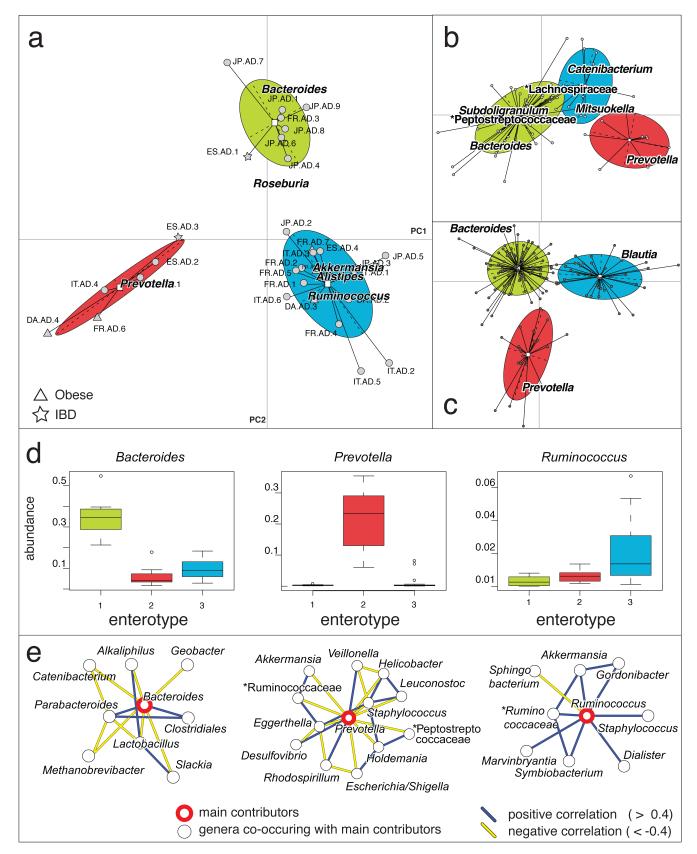
Application in bioinformatics
Connection with linear regression (Optional)
Generalized linear model
Limitations of ordinary linear regression
- Count data (Multinomial/Poisson)
- Dichotomous or Multinomial outcomes (Bernoulli)
- Proportions (values between 0 and 1)
A generalized linear model (GLM) can model different kinds of data.
Components of GLM
- Random components
- Identify \(Y\) and its distribution
- Systematic components
- Linear predictor \(\alpha + \beta_1 x_1 + \dots + \beta_k x_k\)
- Link function
- Given \(\mu=E(Y)\), the link function \(g(\cdot)\) satisfies \[g(\mu)=\alpha+\beta_1x_1+\dots+\beta_kx_k\]
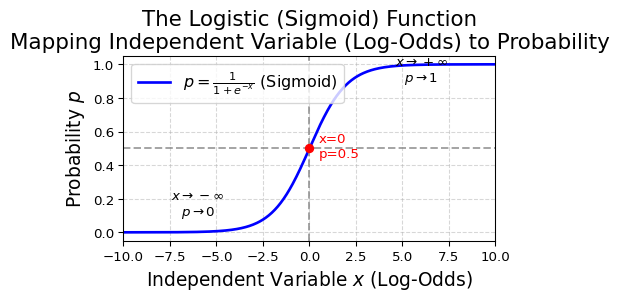
Logistic regression
In logistic regression (binomial outcomes), the link function is
Logit link function
\[
\log\frac{p(x)}{1-p(x)}=\alpha+\beta x
\]
Transform them back to get \(p(x)\)
\[ p(x)=\frac{e^{\alpha+\beta x}}{1+e^{\alpha+\beta x}} \]
Discussion: if we model \(p(x)=\alpha+\beta x\), is it causing any trouble?
- Potential estimate for \(p(x)>1\) or \(p(x)<0\)
Multinomial logistic regression
- How do we compare multiple categories?
- 1 vs. n
- n vs. n
- There are \(k(k-1)/2\) pairs of comparisons, but actually, you only need \(k-1\) non-redundant comparisons.
Let the category \(K\) as the baseline, for each comparison of \(k = 1,\dots,k-1\)
Baseline-category logit model are \[ \log(\frac{p_k(x)}{p_K(x)})=\alpha_k+\beta_kx \quad k=1,\dots,K-1 \]
Interpretation of \(\beta\)
The model assumes \(\ln(\frac{p(x)}{1-p(x)})=\alpha+\beta x\), so for a given change \(\Delta x\),
The odds at \(x + \Delta x\) is
\[ \frac{p(x + \Delta x)}{1-p(x + \Delta x)}=\exp(\alpha+\beta(x+\Delta x)) = \exp(\alpha+\beta(x))\exp(\beta\Delta x) \]
The odds ratio (OR) for \(x\) is
\[ \text{OR}=\frac{\text{odds at}(x+\Delta x)}{\text{odds at}(x)}=\frac{\exp(\alpha+\beta(x+\Delta x))}{\exp(\alpha+\beta(x))} = \exp(\beta\Delta x) \]
- \(\beta\) is the change in the log odds when x increases by 1 unit.
- Confidence interval of OR is not symmetric to the mean.
How about alpha ?
Connections with contingency tables
Logistic regression v.s. Contingency table
| Logistic regression | Contingency table | |
|---|---|---|
| Estimating | OR | OR |
| Test on esstimation | t-test/\(\chi^2\) test | Fisher’s exact test/\(\chi^2\) test |
- Advantage of logistic regression
- Controlling other confounders
- Estimating OR for continuous variables
- Compatible with multinomial outcomes
Back to the kidney stone example
In clinical practice, for stone diameters less than 2cm, doctors usually conduct PCNL, while open surgery is used for stone diameters of 2cm or greater.
| Stone size | Treatment | Success | Fail |
|---|---|---|---|
| Small | OS | 81 | 6 |
| Small | PCNL | 234 | 36 |
| Large | OS | 192 | 71 |
| Large | PCNL | 55 | 25 |
Model: (Success/Fail) ~ Stone size + Treatment
-- Log Odds Ratios & CI w/o Control Stone Size --
2.5% 97.5% Odds Ratio
Intercept 1.28 1.83 1.56
Treatment_OS -0.66 0.08 -0.29
-- Log Odds Ratios & CI with Control Stone Size --
2.5% 97.5% Odds Ratio
Intercept 1.60 2.27 1.94
Stone_Size_Large -1.73 -0.79 -1.26
Treatment_OS -0.09 0.81 0.36Connections to continuous variables
Ordinal logistic regression
For ordinal data (e.g., a scale with scores “No”, “Neutral”, and “Yes”), the following rules apply:
\[ P(x \leq k)=p(x=1) + \dots + p(x=k), \text{where } p(x\leq K) = 1 \]
Intuitively, we can think that the regression is measuring the same thing, but the cutoff for making decisions is different. So, we only need to estimate the cumulative logits:
\[ \log(\frac{p(x\leq j)}{p(x > j)}) =\alpha_j+\beta x \]
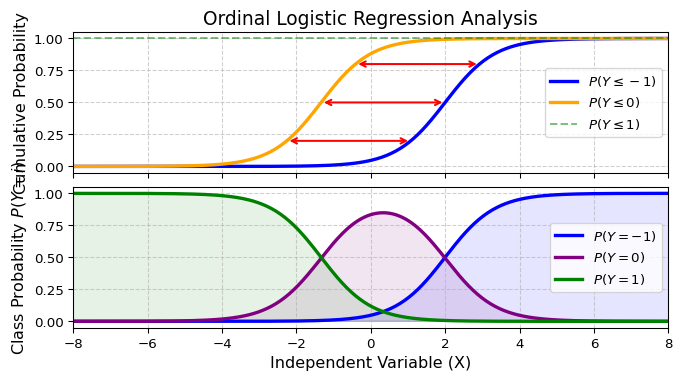
Ordinal logistic regression
Illustration: \(\alpha_{-1}=-3,\alpha_{0}=2, \beta=1.5\)
Implications
- Finding cut-offs for different ordinal outcomes
- If we extend to really continuous case, then we do not need that cut-offs to specific class